Chapter 8
# Install and load packages ---------------
packages <- c(
"tidyverse",
"ggrepel",
"WDI",
"forcats",
"grid",
"gridExtra",
"countrycode",
"lmtest",
"sandwich",
"scales",
"haven",
"kableExtra",
"magrittr",
"rstan"
)
# Change to install = TRUE to install the required packages
pacman::p_load(packages, character.only = TRUE)The stan files used in this document are almost entirely based on the codes written by Lee, Feller, and Rabe-Hesketh. Their original code is found here. Their documentation on this code is found here.
df <- read_dta("http://www.nber.org/~rdehejia/data/nsw_dw.dta") %>%
mutate(
earn74 = re74 / 1000,
earn75 = re75 / 1000,
earn78 = re78 / 1000,
earn74_0 = (re74 == 0),
earn75_0 = (re75 == 0),
earn78_0 = (re78 == 0),
)
cbind(
c(
"age", "education", "married", "nodegree", "black",
"earn'74", "earn'74 = 0", "earn'75", "earn'75 = 0", "earn'78", "earn'78 = 0"
),
df %>%
summarise_at(
vars(age, education, married, nodegree, black, earn74, earn74_0, earn75, earn75_0, earn78, earn78_0),
list(mean, sd)
) %>%
pivot_longer(cols = everything(), names_to = c(".value", "fn"), names_patter = "(.*)(_.*$)") %>%
mutate(
across(-fn, ~ formatC(., digits = 2, format = "f")),
across(-fn, ~ ifelse(fn == "_fn2", paste0("(", ., ")"), .)),
) %>%
select(-fn) %>%
t(),
df %>%
filter(treat == 0) %>%
summarise_at(
vars(age, education, married, nodegree, black, earn74, earn74_0, earn75, earn75_0, earn78, earn78_0),
mean
) %>%
mutate(
across(everything(), ~ formatC(., digits = 2, format = "f")),
) %>%
t(),
df %>%
filter(treat == 1) %>%
summarise_at(
vars(age, education, married, nodegree, black, earn74, earn74_0, earn75, earn75_0, earn78, earn78_0),
mean
) %>%
mutate(
across(everything(), ~ formatC(., digits = 2, format = "f")),
) %>%
t()
) %>%
set_rownames(NULL) %>%
kable("html", booktabs = TRUE, align = c("l", rep("c", 4))) %>%
add_header_above(
c(" ", " ", " ", "($N_c$ = 260)", "($N_t$ = 185)")
) %>%
add_header_above(
c("Covariate", "Mean", "(S.D.)", "Average Controls", "Average Treated")
) %>%
kable_styling(position = "center")| age | 25.37 | (7.10) | 25.05 | 25.82 |
| education | 10.20 | (1.79) | 10.09 | 10.35 |
| married | 0.17 | (0.37) | 0.15 | 0.19 |
| nodegree | 0.78 | (0.41) | 0.83 | 0.71 |
| black | 0.83 | (0.37) | 0.83 | 0.84 |
| earn’74 | 2.10 | (5.36) | 2.11 | 2.10 |
| earn’74 = 0 | 0.73 | (0.44) | 0.75 | 0.71 |
| earn’75 | 1.38 | (3.15) | 1.27 | 1.53 |
| earn’75 = 0 | 0.65 | (0.48) | 0.68 | 0.60 |
| earn’78 | 5.30 | (6.63) | 4.55 | 6.35 |
| earn’78 = 0 | 0.31 | (0.46) | 0.35 | 0.24 |
df %>%
filter(treat == 0) %>%
ggplot(aes(x = earn78)) +
stat_bin(aes(y = ..count../sum(..count..)), binwidth = 1, boundary = 0) +
xlim(c(0, 80)) +
ylim(c(0, 0.5)) +
theme_minimal()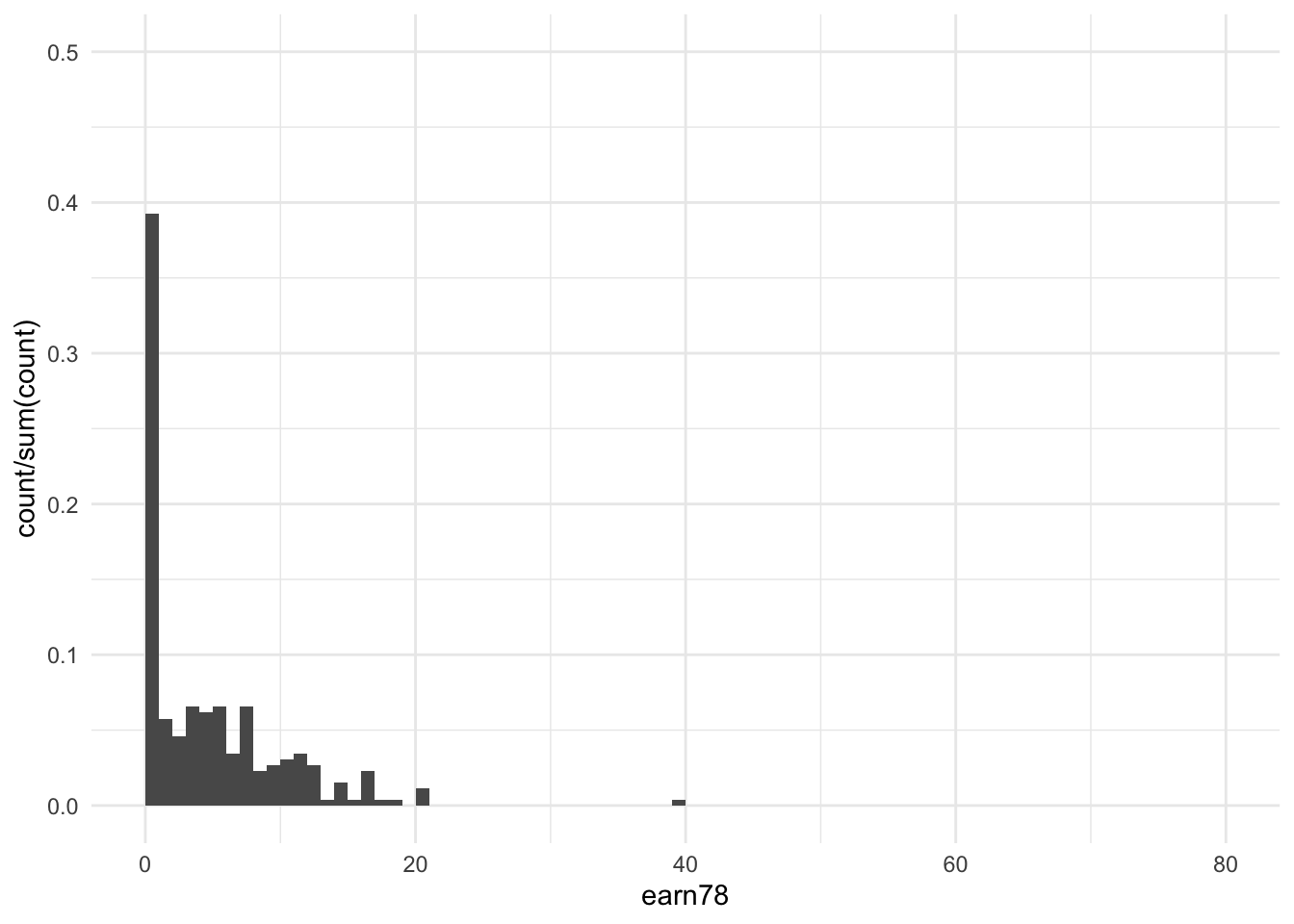
df %>%
filter(treat == 1) %>%
ggplot(aes(x = earn78)) +
stat_bin(aes(y = ..count../sum(..count..)), binwidth = 1, boundary = 0) +
xlim(c(0, 80)) +
ylim(c(0, 0.5)) +
theme_minimal()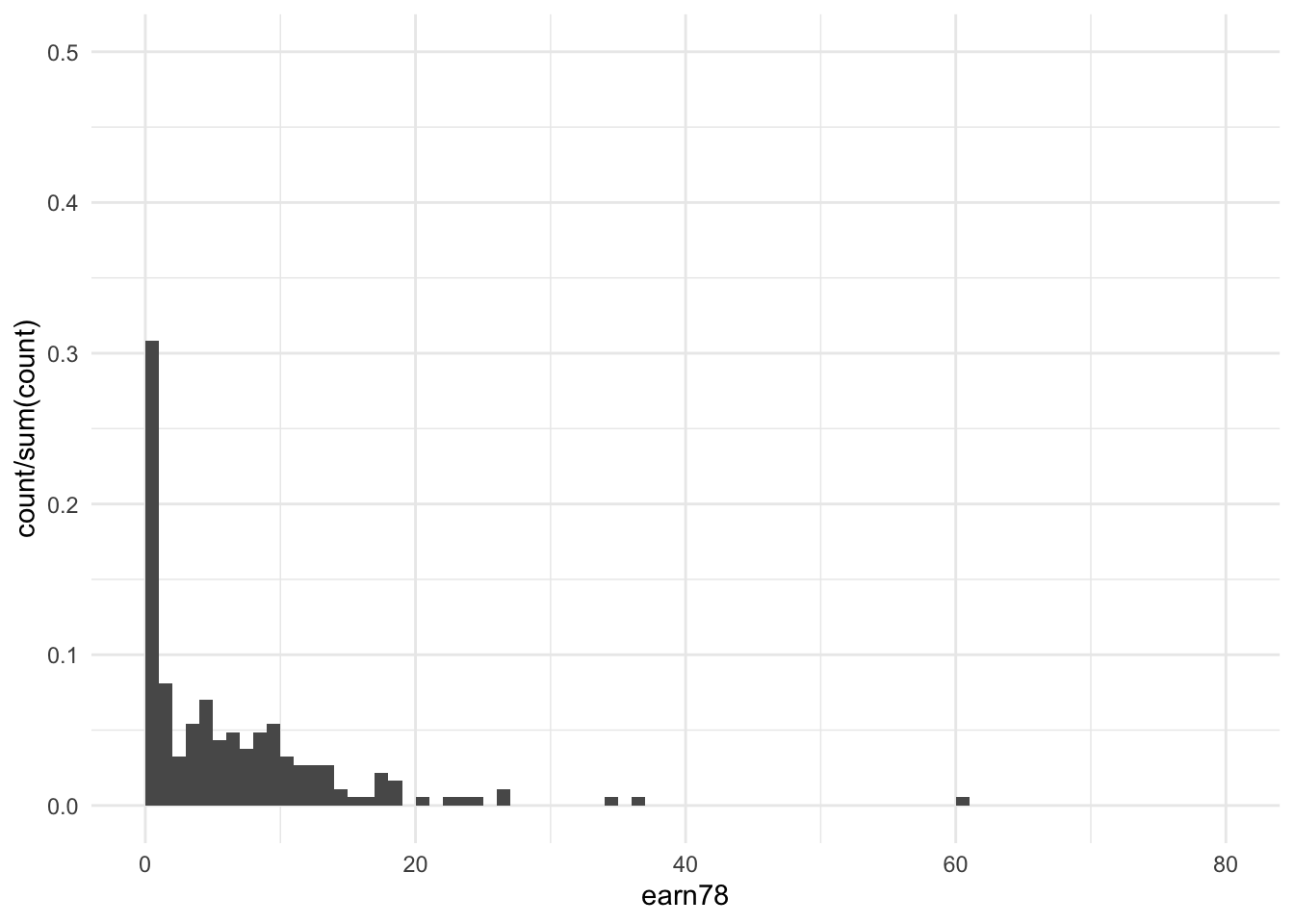
Model 1
stan_data <- list(N = nrow(df), y = df$earn78, w = df$treat)
# Compile and run the stan model
fit_mod1 <- stan(
file = "stan_files/Ch8_Model_01.stan",
data = stan_data,
iter = 1000, chains = 4
)Model 2
stan_data <- list(N = nrow(df), y = df$earn78, w = df$treat)
# Compile and run the stan model
fit_mod2 <- stan(
file = "stan_files/Ch8_Model_02.stan",
data = stan_data,
iter = 1000, chains = 4
)## When you compile models, you are also contributing to development of the NEXT
## Stan compiler. In this version of rstan, we compile your model as usual, but
## also test our new compiler on your syntactically correct model. In this case,
## the new compiler did not work like we hoped. By filing an issue at
## https://github.com/stan-dev/stanc3/issues with your model
## or a minimal example that shows this warning you will be contributing
## valuable information to Stan and ensuring your models continue working. Thank you!
## This message can be avoided by wrapping your function call inside suppressMessages()
## or by first calling rstan_options(javascript = FALSE).
## Semantic error in 'string', line 5, column 7 to column 15:
## Identifier 'quantile' clashes with Stan Math library function.##
## SAMPLING FOR MODEL 'Ch8_Model_02' NOW (CHAIN 1).
## Chain 1:
## Chain 1: Gradient evaluation took 7.1e-05 seconds
## Chain 1: 1000 transitions using 10 leapfrog steps per transition would take 0.71 seconds.
## Chain 1: Adjust your expectations accordingly!
## Chain 1:
## Chain 1:
## Chain 1: Iteration: 1 / 1000 [ 0%] (Warmup)
## Chain 1: Iteration: 100 / 1000 [ 10%] (Warmup)
## Chain 1: Iteration: 200 / 1000 [ 20%] (Warmup)
## Chain 1: Iteration: 300 / 1000 [ 30%] (Warmup)
## Chain 1: Iteration: 400 / 1000 [ 40%] (Warmup)
## Chain 1: Iteration: 500 / 1000 [ 50%] (Warmup)
## Chain 1: Iteration: 501 / 1000 [ 50%] (Sampling)
## Chain 1: Iteration: 600 / 1000 [ 60%] (Sampling)
## Chain 1: Iteration: 700 / 1000 [ 70%] (Sampling)
## Chain 1: Iteration: 800 / 1000 [ 80%] (Sampling)
## Chain 1: Iteration: 900 / 1000 [ 90%] (Sampling)
## Chain 1: Iteration: 1000 / 1000 [100%] (Sampling)
## Chain 1:
## Chain 1: Elapsed Time: 0.231515 seconds (Warm-up)
## Chain 1: 0.169078 seconds (Sampling)
## Chain 1: 0.400593 seconds (Total)
## Chain 1:
##
## SAMPLING FOR MODEL 'Ch8_Model_02' NOW (CHAIN 2).
## Chain 2:
## Chain 2: Gradient evaluation took 3.9e-05 seconds
## Chain 2: 1000 transitions using 10 leapfrog steps per transition would take 0.39 seconds.
## Chain 2: Adjust your expectations accordingly!
## Chain 2:
## Chain 2:
## Chain 2: Iteration: 1 / 1000 [ 0%] (Warmup)
## Chain 2: Iteration: 100 / 1000 [ 10%] (Warmup)
## Chain 2: Iteration: 200 / 1000 [ 20%] (Warmup)
## Chain 2: Iteration: 300 / 1000 [ 30%] (Warmup)
## Chain 2: Iteration: 400 / 1000 [ 40%] (Warmup)
## Chain 2: Iteration: 500 / 1000 [ 50%] (Warmup)
## Chain 2: Iteration: 501 / 1000 [ 50%] (Sampling)
## Chain 2: Iteration: 600 / 1000 [ 60%] (Sampling)
## Chain 2: Iteration: 700 / 1000 [ 70%] (Sampling)
## Chain 2: Iteration: 800 / 1000 [ 80%] (Sampling)
## Chain 2: Iteration: 900 / 1000 [ 90%] (Sampling)
## Chain 2: Iteration: 1000 / 1000 [100%] (Sampling)
## Chain 2:
## Chain 2: Elapsed Time: 0.222419 seconds (Warm-up)
## Chain 2: 0.178699 seconds (Sampling)
## Chain 2: 0.401118 seconds (Total)
## Chain 2:
##
## SAMPLING FOR MODEL 'Ch8_Model_02' NOW (CHAIN 3).
## Chain 3:
## Chain 3: Gradient evaluation took 3.6e-05 seconds
## Chain 3: 1000 transitions using 10 leapfrog steps per transition would take 0.36 seconds.
## Chain 3: Adjust your expectations accordingly!
## Chain 3:
## Chain 3:
## Chain 3: Iteration: 1 / 1000 [ 0%] (Warmup)
## Chain 3: Iteration: 100 / 1000 [ 10%] (Warmup)
## Chain 3: Iteration: 200 / 1000 [ 20%] (Warmup)
## Chain 3: Iteration: 300 / 1000 [ 30%] (Warmup)
## Chain 3: Iteration: 400 / 1000 [ 40%] (Warmup)
## Chain 3: Iteration: 500 / 1000 [ 50%] (Warmup)
## Chain 3: Iteration: 501 / 1000 [ 50%] (Sampling)
## Chain 3: Iteration: 600 / 1000 [ 60%] (Sampling)
## Chain 3: Iteration: 700 / 1000 [ 70%] (Sampling)
## Chain 3: Iteration: 800 / 1000 [ 80%] (Sampling)
## Chain 3: Iteration: 900 / 1000 [ 90%] (Sampling)
## Chain 3: Iteration: 1000 / 1000 [100%] (Sampling)
## Chain 3:
## Chain 3: Elapsed Time: 0.21897 seconds (Warm-up)
## Chain 3: 0.167248 seconds (Sampling)
## Chain 3: 0.386218 seconds (Total)
## Chain 3:
##
## SAMPLING FOR MODEL 'Ch8_Model_02' NOW (CHAIN 4).
## Chain 4:
## Chain 4: Gradient evaluation took 3.5e-05 seconds
## Chain 4: 1000 transitions using 10 leapfrog steps per transition would take 0.35 seconds.
## Chain 4: Adjust your expectations accordingly!
## Chain 4:
## Chain 4:
## Chain 4: Iteration: 1 / 1000 [ 0%] (Warmup)
## Chain 4: Iteration: 100 / 1000 [ 10%] (Warmup)
## Chain 4: Iteration: 200 / 1000 [ 20%] (Warmup)
## Chain 4: Iteration: 300 / 1000 [ 30%] (Warmup)
## Chain 4: Iteration: 400 / 1000 [ 40%] (Warmup)
## Chain 4: Iteration: 500 / 1000 [ 50%] (Warmup)
## Chain 4: Iteration: 501 / 1000 [ 50%] (Sampling)
## Chain 4: Iteration: 600 / 1000 [ 60%] (Sampling)
## Chain 4: Iteration: 700 / 1000 [ 70%] (Sampling)
## Chain 4: Iteration: 800 / 1000 [ 80%] (Sampling)
## Chain 4: Iteration: 900 / 1000 [ 90%] (Sampling)
## Chain 4: Iteration: 1000 / 1000 [100%] (Sampling)
## Chain 4:
## Chain 4: Elapsed Time: 0.218884 seconds (Warm-up)
## Chain 4: 0.179702 seconds (Sampling)
## Chain 4: 0.398586 seconds (Total)
## Chain 4:Model 3
# Add nine mean-centered covariates and their interaction terms
x <- as.matrix(df %>% select(age, education, married, nodegree, black, earn74, earn74_0, earn75, earn75_0))
x_mean_mat <- matrix(rep(apply(x, 2, mean, na.rm = TRUE), each = nrow(x)), nrow = nrow(x))
x_c_mat <- x - x_mean_mat
xw_inter <- x_c_mat * df$treat
colnames(xw_inter) <- paste0(colnames(x), "_w")
# Collect data into a list format suitable for Stan
stan_data <- list(
N = nrow(df),
N_cov = ncol(x_c_mat),
y = df$earn78,
w = df$treat,
x = x_c_mat,
xw_inter = xw_inter
)
# Compile and run the stan model
fit_mod3 <- stan(
file = "stan_files/Ch8_Model_03.stan",
data = stan_data,
iter = 1000, chains = 4
)## When you compile models, you are also contributing to development of the NEXT
## Stan compiler. In this version of rstan, we compile your model as usual, but
## also test our new compiler on your syntactically correct model. In this case,
## the new compiler did not work like we hoped. By filing an issue at
## https://github.com/stan-dev/stanc3/issues with your model
## or a minimal example that shows this warning you will be contributing
## valuable information to Stan and ensuring your models continue working. Thank you!
## This message can be avoided by wrapping your function call inside suppressMessages()
## or by first calling rstan_options(javascript = FALSE).
## Semantic error in 'string', line 5, column 7 to column 15:
## Identifier 'quantile' clashes with Stan Math library function.##
## SAMPLING FOR MODEL 'Ch8_Model_03' NOW (CHAIN 1).
## Chain 1:
## Chain 1: Gradient evaluation took 0.000156 seconds
## Chain 1: 1000 transitions using 10 leapfrog steps per transition would take 1.56 seconds.
## Chain 1: Adjust your expectations accordingly!
## Chain 1:
## Chain 1:
## Chain 1: Iteration: 1 / 1000 [ 0%] (Warmup)
## Chain 1: Iteration: 100 / 1000 [ 10%] (Warmup)
## Chain 1: Iteration: 200 / 1000 [ 20%] (Warmup)
## Chain 1: Iteration: 300 / 1000 [ 30%] (Warmup)
## Chain 1: Iteration: 400 / 1000 [ 40%] (Warmup)
## Chain 1: Iteration: 500 / 1000 [ 50%] (Warmup)
## Chain 1: Iteration: 501 / 1000 [ 50%] (Sampling)
## Chain 1: Iteration: 600 / 1000 [ 60%] (Sampling)
## Chain 1: Iteration: 700 / 1000 [ 70%] (Sampling)
## Chain 1: Iteration: 800 / 1000 [ 80%] (Sampling)
## Chain 1: Iteration: 900 / 1000 [ 90%] (Sampling)
## Chain 1: Iteration: 1000 / 1000 [100%] (Sampling)
## Chain 1:
## Chain 1: Elapsed Time: 1.5093 seconds (Warm-up)
## Chain 1: 0.625582 seconds (Sampling)
## Chain 1: 2.13488 seconds (Total)
## Chain 1:
##
## SAMPLING FOR MODEL 'Ch8_Model_03' NOW (CHAIN 2).
## Chain 2:
## Chain 2: Gradient evaluation took 6.6e-05 seconds
## Chain 2: 1000 transitions using 10 leapfrog steps per transition would take 0.66 seconds.
## Chain 2: Adjust your expectations accordingly!
## Chain 2:
## Chain 2:
## Chain 2: Iteration: 1 / 1000 [ 0%] (Warmup)
## Chain 2: Iteration: 100 / 1000 [ 10%] (Warmup)
## Chain 2: Iteration: 200 / 1000 [ 20%] (Warmup)
## Chain 2: Iteration: 300 / 1000 [ 30%] (Warmup)
## Chain 2: Iteration: 400 / 1000 [ 40%] (Warmup)
## Chain 2: Iteration: 500 / 1000 [ 50%] (Warmup)
## Chain 2: Iteration: 501 / 1000 [ 50%] (Sampling)
## Chain 2: Iteration: 600 / 1000 [ 60%] (Sampling)
## Chain 2: Iteration: 700 / 1000 [ 70%] (Sampling)
## Chain 2: Iteration: 800 / 1000 [ 80%] (Sampling)
## Chain 2: Iteration: 900 / 1000 [ 90%] (Sampling)
## Chain 2: Iteration: 1000 / 1000 [100%] (Sampling)
## Chain 2:
## Chain 2: Elapsed Time: 1.48241 seconds (Warm-up)
## Chain 2: 0.707962 seconds (Sampling)
## Chain 2: 2.19037 seconds (Total)
## Chain 2:
##
## SAMPLING FOR MODEL 'Ch8_Model_03' NOW (CHAIN 3).
## Chain 3:
## Chain 3: Gradient evaluation took 6.5e-05 seconds
## Chain 3: 1000 transitions using 10 leapfrog steps per transition would take 0.65 seconds.
## Chain 3: Adjust your expectations accordingly!
## Chain 3:
## Chain 3:
## Chain 3: Iteration: 1 / 1000 [ 0%] (Warmup)
## Chain 3: Iteration: 100 / 1000 [ 10%] (Warmup)
## Chain 3: Iteration: 200 / 1000 [ 20%] (Warmup)
## Chain 3: Iteration: 300 / 1000 [ 30%] (Warmup)
## Chain 3: Iteration: 400 / 1000 [ 40%] (Warmup)
## Chain 3: Iteration: 500 / 1000 [ 50%] (Warmup)
## Chain 3: Iteration: 501 / 1000 [ 50%] (Sampling)
## Chain 3: Iteration: 600 / 1000 [ 60%] (Sampling)
## Chain 3: Iteration: 700 / 1000 [ 70%] (Sampling)
## Chain 3: Iteration: 800 / 1000 [ 80%] (Sampling)
## Chain 3: Iteration: 900 / 1000 [ 90%] (Sampling)
## Chain 3: Iteration: 1000 / 1000 [100%] (Sampling)
## Chain 3:
## Chain 3: Elapsed Time: 1.52593 seconds (Warm-up)
## Chain 3: 0.603981 seconds (Sampling)
## Chain 3: 2.12991 seconds (Total)
## Chain 3:
##
## SAMPLING FOR MODEL 'Ch8_Model_03' NOW (CHAIN 4).
## Chain 4:
## Chain 4: Gradient evaluation took 6e-05 seconds
## Chain 4: 1000 transitions using 10 leapfrog steps per transition would take 0.6 seconds.
## Chain 4: Adjust your expectations accordingly!
## Chain 4:
## Chain 4:
## Chain 4: Iteration: 1 / 1000 [ 0%] (Warmup)
## Chain 4: Iteration: 100 / 1000 [ 10%] (Warmup)
## Chain 4: Iteration: 200 / 1000 [ 20%] (Warmup)
## Chain 4: Iteration: 300 / 1000 [ 30%] (Warmup)
## Chain 4: Iteration: 400 / 1000 [ 40%] (Warmup)
## Chain 4: Iteration: 500 / 1000 [ 50%] (Warmup)
## Chain 4: Iteration: 501 / 1000 [ 50%] (Sampling)
## Chain 4: Iteration: 600 / 1000 [ 60%] (Sampling)
## Chain 4: Iteration: 700 / 1000 [ 70%] (Sampling)
## Chain 4: Iteration: 800 / 1000 [ 80%] (Sampling)
## Chain 4: Iteration: 900 / 1000 [ 90%] (Sampling)
## Chain 4: Iteration: 1000 / 1000 [100%] (Sampling)
## Chain 4:
## Chain 4: Elapsed Time: 1.5719 seconds (Warm-up)
## Chain 4: 0.583757 seconds (Sampling)
## Chain 4: 2.15565 seconds (Total)
## Chain 4:Model 4
# Add nine mean-centered covariates and their interaction terms
x <- as.matrix(
df %>%
select(age, education, married, nodegree, black, earn74, earn74_0, earn75, earn75_0) %>%
mutate(
earn74_0 = (earn74 > 0),
earn75_0 = (earn75 > 0),
)
)
x_mean_mat <- matrix(rep(apply(x, 2, mean, na.rm = TRUE), each = nrow(x)), nrow = nrow(x))
x_c_mat <- x - x_mean_mat
xw_inter <- x_c_mat * df$treat
colnames(xw_inter) <- paste0(colnames(x), "_w")
# Collect data into a list format suitable for Stan
stan_data <- list(
N = nrow(df),
N_cov = ncol(x_c_mat),
y = df$earn78,
y_pos = as.numeric(df$earn78 > 0),
z = df$treat,
x = x_c_mat,
xz_inter = xw_inter,
# x = x,
# xz_inter = x * df$treat,
rho = 0
)
# Compile and run the stan model
fit_mod4 <- stan(
file = "stan_files/Ch8_Model_04.stan",
data = stan_data,
iter = 1000, chains = 4
)## When you compile models, you are also contributing to development of the NEXT
## Stan compiler. In this version of rstan, we compile your model as usual, but
## also test our new compiler on your syntactically correct model. In this case,
## the new compiler did not work like we hoped. By filing an issue at
## https://github.com/stan-dev/stanc3/issues with your model
## or a minimal example that shows this warning you will be contributing
## valuable information to Stan and ensuring your models continue working. Thank you!
## This message can be avoided by wrapping your function call inside suppressMessages()
## or by first calling rstan_options(javascript = FALSE).
## Semantic error in 'string', line 2, column 7 to column 15:
## Identifier 'quantile' clashes with Stan Math library function.##
## SAMPLING FOR MODEL 'Ch8_Model_04' NOW (CHAIN 1).
## Chain 1: Rejecting initial value:
## Chain 1: Error evaluating the log probability at the initial value.
## Chain 1: Exception: lognormal_lpdf: Scale parameter is -1.90969, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 1: Rejecting initial value:
## Chain 1: Error evaluating the log probability at the initial value.
## Chain 1: Exception: lognormal_lpdf: Scale parameter is -0.494532, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 1: Rejecting initial value:
## Chain 1: Error evaluating the log probability at the initial value.
## Chain 1: Exception: lognormal_lpdf: Scale parameter is -0.72984, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 1: Rejecting initial value:
## Chain 1: Error evaluating the log probability at the initial value.
## Chain 1: Exception: lognormal_lpdf: Scale parameter is -0.360274, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 1: Rejecting initial value:
## Chain 1: Error evaluating the log probability at the initial value.
## Chain 1: Exception: lognormal_lpdf: Scale parameter is -0.26098, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 1: Rejecting initial value:
## Chain 1: Error evaluating the log probability at the initial value.
## Chain 1: Exception: lognormal_lpdf: Scale parameter is -0.678968, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 1: Rejecting initial value:
## Chain 1: Error evaluating the log probability at the initial value.
## Chain 1: Exception: lognormal_lpdf: Scale parameter is -0.877445, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 1: Rejecting initial value:
## Chain 1: Error evaluating the log probability at the initial value.
## Chain 1: Exception: lognormal_lpdf: Scale parameter is -1.93821, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 1: Rejecting initial value:
## Chain 1: Error evaluating the log probability at the initial value.
## Chain 1: Exception: lognormal_lpdf: Scale parameter is -0.00674907, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 1: Rejecting initial value:
## Chain 1: Error evaluating the log probability at the initial value.
## Chain 1: Exception: lognormal_lpdf: Scale parameter is -1.00088, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 1:
## Chain 1: Gradient evaluation took 0.000309 seconds
## Chain 1: 1000 transitions using 10 leapfrog steps per transition would take 3.09 seconds.
## Chain 1: Adjust your expectations accordingly!
## Chain 1:
## Chain 1:
## Chain 1: Iteration: 1 / 1000 [ 0%] (Warmup)
## Chain 1: Iteration: 100 / 1000 [ 10%] (Warmup)
## Chain 1: Iteration: 200 / 1000 [ 20%] (Warmup)
## Chain 1: Iteration: 300 / 1000 [ 30%] (Warmup)
## Chain 1: Iteration: 400 / 1000 [ 40%] (Warmup)
## Chain 1: Iteration: 500 / 1000 [ 50%] (Warmup)
## Chain 1: Iteration: 501 / 1000 [ 50%] (Sampling)
## Chain 1: Iteration: 600 / 1000 [ 60%] (Sampling)
## Chain 1: Iteration: 700 / 1000 [ 70%] (Sampling)
## Chain 1: Iteration: 800 / 1000 [ 80%] (Sampling)
## Chain 1: Iteration: 900 / 1000 [ 90%] (Sampling)
## Chain 1: Iteration: 1000 / 1000 [100%] (Sampling)
## Chain 1:
## Chain 1: Elapsed Time: 9.02484 seconds (Warm-up)
## Chain 1: 3.19613 seconds (Sampling)
## Chain 1: 12.221 seconds (Total)
## Chain 1:
##
## SAMPLING FOR MODEL 'Ch8_Model_04' NOW (CHAIN 2).
## Chain 2: Rejecting initial value:
## Chain 2: Error evaluating the log probability at the initial value.
## Chain 2: Exception: lognormal_lpdf: Scale parameter is -0.0542545, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 2: Rejecting initial value:
## Chain 2: Error evaluating the log probability at the initial value.
## Chain 2: Exception: lognormal_lpdf: Scale parameter is -0.269141, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 2: Rejecting initial value:
## Chain 2: Error evaluating the log probability at the initial value.
## Chain 2: Exception: lognormal_lpdf: Scale parameter is -1.19248, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 2: Rejecting initial value:
## Chain 2: Error evaluating the log probability at the initial value.
## Chain 2: Exception: lognormal_lpdf: Scale parameter is -1.18821, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 2: Rejecting initial value:
## Chain 2: Error evaluating the log probability at the initial value.
## Chain 2: Exception: lognormal_lpdf: Scale parameter is -1.32215, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 2:
## Chain 2: Gradient evaluation took 0.0002 seconds
## Chain 2: 1000 transitions using 10 leapfrog steps per transition would take 2 seconds.
## Chain 2: Adjust your expectations accordingly!
## Chain 2:
## Chain 2:
## Chain 2: Iteration: 1 / 1000 [ 0%] (Warmup)
## Chain 2: Iteration: 100 / 1000 [ 10%] (Warmup)
## Chain 2: Iteration: 200 / 1000 [ 20%] (Warmup)
## Chain 2: Iteration: 300 / 1000 [ 30%] (Warmup)
## Chain 2: Iteration: 400 / 1000 [ 40%] (Warmup)
## Chain 2: Iteration: 500 / 1000 [ 50%] (Warmup)
## Chain 2: Iteration: 501 / 1000 [ 50%] (Sampling)
## Chain 2: Iteration: 600 / 1000 [ 60%] (Sampling)
## Chain 2: Iteration: 700 / 1000 [ 70%] (Sampling)
## Chain 2: Iteration: 800 / 1000 [ 80%] (Sampling)
## Chain 2: Iteration: 900 / 1000 [ 90%] (Sampling)
## Chain 2: Iteration: 1000 / 1000 [100%] (Sampling)
## Chain 2:
## Chain 2: Elapsed Time: 8.27101 seconds (Warm-up)
## Chain 2: 2.94859 seconds (Sampling)
## Chain 2: 11.2196 seconds (Total)
## Chain 2:
##
## SAMPLING FOR MODEL 'Ch8_Model_04' NOW (CHAIN 3).
## Chain 3:
## Chain 3: Gradient evaluation took 0.000197 seconds
## Chain 3: 1000 transitions using 10 leapfrog steps per transition would take 1.97 seconds.
## Chain 3: Adjust your expectations accordingly!
## Chain 3:
## Chain 3:
## Chain 3: Iteration: 1 / 1000 [ 0%] (Warmup)
## Chain 3: Iteration: 100 / 1000 [ 10%] (Warmup)
## Chain 3: Iteration: 200 / 1000 [ 20%] (Warmup)
## Chain 3: Iteration: 300 / 1000 [ 30%] (Warmup)
## Chain 3: Iteration: 400 / 1000 [ 40%] (Warmup)
## Chain 3: Iteration: 500 / 1000 [ 50%] (Warmup)
## Chain 3: Iteration: 501 / 1000 [ 50%] (Sampling)
## Chain 3: Iteration: 600 / 1000 [ 60%] (Sampling)
## Chain 3: Iteration: 700 / 1000 [ 70%] (Sampling)
## Chain 3: Iteration: 800 / 1000 [ 80%] (Sampling)
## Chain 3: Iteration: 900 / 1000 [ 90%] (Sampling)
## Chain 3: Iteration: 1000 / 1000 [100%] (Sampling)
## Chain 3:
## Chain 3: Elapsed Time: 7.75415 seconds (Warm-up)
## Chain 3: 2.80312 seconds (Sampling)
## Chain 3: 10.5573 seconds (Total)
## Chain 3:
##
## SAMPLING FOR MODEL 'Ch8_Model_04' NOW (CHAIN 4).
## Chain 4: Rejecting initial value:
## Chain 4: Error evaluating the log probability at the initial value.
## Chain 4: Exception: lognormal_lpdf: Scale parameter is -1.52753, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 4: Rejecting initial value:
## Chain 4: Error evaluating the log probability at the initial value.
## Chain 4: Exception: lognormal_lpdf: Scale parameter is -0.500182, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 4: Rejecting initial value:
## Chain 4: Error evaluating the log probability at the initial value.
## Chain 4: Exception: lognormal_lpdf: Scale parameter is -0.0603489, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 4: Rejecting initial value:
## Chain 4: Error evaluating the log probability at the initial value.
## Chain 4: Exception: lognormal_lpdf: Scale parameter is -1.84628, but must be > 0! (in 'modelc8e2427df18_Ch8_Model_04' at line 78)
##
## Chain 4:
## Chain 4: Gradient evaluation took 0.000195 seconds
## Chain 4: 1000 transitions using 10 leapfrog steps per transition would take 1.95 seconds.
## Chain 4: Adjust your expectations accordingly!
## Chain 4:
## Chain 4:
## Chain 4: Iteration: 1 / 1000 [ 0%] (Warmup)
## Chain 4: Iteration: 100 / 1000 [ 10%] (Warmup)
## Chain 4: Iteration: 200 / 1000 [ 20%] (Warmup)
## Chain 4: Iteration: 300 / 1000 [ 30%] (Warmup)
## Chain 4: Iteration: 400 / 1000 [ 40%] (Warmup)
## Chain 4: Iteration: 500 / 1000 [ 50%] (Warmup)
## Chain 4: Iteration: 501 / 1000 [ 50%] (Sampling)
## Chain 4: Iteration: 600 / 1000 [ 60%] (Sampling)
## Chain 4: Iteration: 700 / 1000 [ 70%] (Sampling)
## Chain 4: Iteration: 800 / 1000 [ 80%] (Sampling)
## Chain 4: Iteration: 900 / 1000 [ 90%] (Sampling)
## Chain 4: Iteration: 1000 / 1000 [100%] (Sampling)
## Chain 4:
## Chain 4: Elapsed Time: 8.54278 seconds (Warm-up)
## Chain 4: 2.96939 seconds (Sampling)
## Chain 4: 11.5122 seconds (Total)
## Chain 4:Table 8.6
param <- c("tau_fs", "tau_qte25", "tau_qte50", "tau_qte75" )
cbind(
c("No", "No", "Yes", "Yes"),
c("No", "Yes", "Yes", "Yes"),
c("No", "Yes", "Yes", "Yes"),
c("No", "No", "No", "Yes"),
map(
list(fit_mod1, fit_mod2, fit_mod3, fit_mod4),
function(fit_mod) {
fit_mod_summary <- summary(fit_mod)$summary[param, c("mean", "sd")]
fit_mod_summary <- cbind(
formatC(fit_mod_summary[,1], digits = 2, format = "f"),
paste0("(", formatC(fit_mod_summary[,2], digits = 2, format = "f"), ")")
)
return(map(seq(nrow(fit_mod_summary)), ~ fit_mod_summary[.,] %>% as.vector) %>% unlist())
}
) %>%
bind_cols %>%
t()
) %>%
set_rownames(NULL) %>%
kable("html", booktabs = TRUE, align = c("l", rep("c", 11))) %>%
add_header_above(
c(
"Mean\nCovariate\nDependent", "Variance\nTreatment\nSpecific",
"Potential\nOutcome\nIndependent", "Two-\nPart\nModel",
rep(c("Mean", "(S.D.)"), 4)
)
) %>%
add_header_above(
c(" ", " ", " ", " ",
"Mean Effect" = 2, "0.25 quant" = 2, "0.50 quant" = 2, "0.75 quant" = 2
)
) %>%
add_header_above(
c(
" ", " ", " ", " ", " ", " ",
"Effects on Quantiles" = 6
)
) %>%
kable_styling(position = "center")## New names:
## * NA -> ...1
## * NA -> ...2
## * NA -> ...3
## * NA -> ...4| No | No | No | No | 1.80 | (0.64) | 1.80 | (0.64) | 1.80 | (0.64) | 1.80 | (0.64) |
| No | Yes | Yes | No | 1.79 | (0.49) | 0.63 | (0.35) | 1.65 | (0.56) | 3.10 | (0.63) |
| Yes | Yes | Yes | No | 1.56 | (0.51) | 0.42 | (0.34) | 1.39 | (0.56) | 2.88 | (0.64) |
| Yes | Yes | Yes | Yes | 1.82 | (0.88) | 0.23 | (0.29) | 1.01 | (0.54) | 1.81 | (0.75) |
Table 8.7
summary(fit_mod4)$summary %>% rownames## [1] "alpha_bin" "beta_bin[1]" "beta_bin[2]" "beta_bin[3]" "beta_bin[4]"
## [6] "beta_bin[5]" "beta_bin[6]" "beta_bin[7]" "beta_bin[8]" "beta_bin[9]"
## [11] "beta_inter_bin[1]" "beta_inter_bin[2]" "beta_inter_bin[3]" "beta_inter_bin[4]" "beta_inter_bin[5]"
## [16] "beta_inter_bin[6]" "beta_inter_bin[7]" "beta_inter_bin[8]" "beta_inter_bin[9]" "tau_bin"
## [21] "alpha_cont" "beta_cont[1]" "beta_cont[2]" "beta_cont[3]" "beta_cont[4]"
## [26] "beta_cont[5]" "beta_cont[6]" "beta_cont[7]" "beta_cont[8]" "beta_cont[9]"
## [31] "beta_inter_cont[1]" "beta_inter_cont[2]" "beta_inter_cont[3]" "beta_inter_cont[4]" "beta_inter_cont[5]"
## [36] "beta_inter_cont[6]" "beta_inter_cont[7]" "beta_inter_cont[8]" "beta_inter_cont[9]" "tau_cont"
## [41] "sigma_t" "sigma_c" "y0_bar" "y1_bar" "tau_fs"
## [46] "log_sigma_c" "log_sigma_t" "tau_qte25" "tau_qte50" "tau_qte75"
## [51] "lp__"param1 <- c("alpha_cont", paste0("beta_cont[", seq(1,9), "]"))
param2 <- c("tau_cont", paste0("beta_inter_cont[", seq(1,9), "]"))
param3 <- c("alpha_bin", paste0("beta_bin[", seq(1,9), "]"))
param4 <- c("tau_bin", paste0("beta_inter_bin[", seq(1,9), "]"))
cbind(
c(
"intercept", "age", "education", "married",
"nodegree", "black", "earn '74", "earn '74 $>$ 0",
"earn '75", "earn '75 $>$ 0"
),
map(
list(param1, param2, param3, param4),
function(param) {
fit_mod_summary <- summary(fit_mod4)$summary[param, c("mean", "sd")]
fit_mod_summary <- cbind(
formatC(fit_mod_summary[,1], digits = 2, format = "f"),
paste0("(", formatC(fit_mod_summary[,2], digits = 2, format = "f"), ")")
)
} %>%
as_tibble()
) %>%
bind_cols %>%
as.matrix()
) %>%
rbind(
c(
"ln($\\sigma_c$)",
summary(fit_mod4)$summary["log_sigma_c", c("mean")] %>% formatC(digits = 3, format = "f"),
paste0("(", summary(fit_mod4)$summary["log_sigma_c", c("sd")] %>% formatC(digits = 3, format = "f"), ")"),
rep(" ", 6)
),
c(
"ln($\\sigma_t$)",
summary(fit_mod4)$summary["log_sigma_t", c("mean")] %>% formatC(digits = 3, format = "f"),
paste0("(", summary(fit_mod4)$summary["log_sigma_t", c("sd")] %>% formatC(digits = 3, format = "f"), ")"),
rep(" ", 6)
)
) %>%
set_colnames(NULL) %>%
kable("html", booktabs = TRUE, align = c("l", rep("c", 8))) %>%
add_header_above(
c(" ", rep(c("Mean", "(S.D.)"), 4))
) %>%
add_header_above(
c("Covariate", "$\\beta_c$" = 2, "$\\beta_t - \\beta_c$" = 2, "$\\gamma_0$" = 2, "$\\gamma_1 - \\gamma_0$" = 2)
) %>%
kable_styling(position = "center")## New names:
## * V1 -> V1...1
## * V2 -> V2...2
## * V1 -> V1...3
## * V2 -> V2...4
## * V1 -> V1...5
## * ...| intercept | 1.58 | (0.08) | 0.07 | (0.13) | 0.67 | (0.14) | 0.78 | (0.28) |
| age | 0.02 | (0.01) | -0.02 | (0.02) | -0.01 | (0.02) | 0.02 | (0.03) |
| education | 0.01 | (0.06) | 0.01 | (0.09) | -0.05 | (0.12) | 0.02 | (0.18) |
| married | -0.23 | (0.25) | 0.36 | (0.36) | -0.17 | (0.39) | 0.91 | (0.71) |
| nodegree | -0.01 | (0.27) | -0.24 | (0.38) | -0.28 | (0.48) | -0.25 | (0.76) |
| black | -0.44 | (0.20) | 0.36 | (0.31) | -1.08 | (0.43) | -0.79 | (1.02) |
| earn ’74 | -0.01 | (0.02) | 0.01 | (0.03) | 0.01 | (0.04) | -0.02 | (0.08) |
| earn ’74 \(>\) 0 | 0.18 | (0.32) | -0.57 | (0.46) | 1.00 | (0.55) | -3.09 | (1.14) |
| earn ’75 | 0.02 | (0.04) | 0.01 | (0.05) | 0.00 | (0.08) | 0.20 | (0.17) |
| earn ’75 \(>\) 0 | -0.06 | (0.29) | 0.18 | (0.40) | -0.61 | (0.46) | 2.15 | (1.07) |
| ln(\(\sigma_c\)) | 0.003 | (0.057) | ||||||
| ln(\(\sigma_t\)) | 0.052 | (0.063) |